About MepAnti
Research Projects
The project will be focussed on the elucidation of crystal structures of constituent enzymes of the MEP pathway from pathogenic organisms (Pseudomonas aeruginosa, Plasmodium falciparum, Mycobacterium tuberculosis, Escherichia coli).
Over the past decades, structural biologists have extensively studied the MEP pathway enzymes. Nevertheless, important gaps remain to be addressed. By adopting an extensive structural-genomics approach for each constituent enzyme, we will be able to shed light on the structural features of the enzymes across a series of important pathogens.
The students will be trained to establish constructs and crystallization conditions and to carry out high-throughput screening of fragment libraries by X-ray crystallography.
 The drug-discovery projects will exploit established as well as innovative hit-identification strategies. Structure- and fragment-based design, classical structure-activity relationship (SAR) study, dynamic combinatorial chemistry and kinetic target-guided synthesis will be used to identify novel, selective and potent inhibitors of the MEP pathway enzymes.
The drug-discovery projects will exploit established as well as innovative hit-identification strategies. Structure- and fragment-based design, classical structure-activity relationship (SAR) study, dynamic combinatorial chemistry and kinetic target-guided synthesis will be used to identify novel, selective and potent inhibitors of the MEP pathway enzymes.
The binding mode of the most promising hits will be determined by saturation-transfer difference (STD)-NMR, site-directed mutagenesis and co-crystal structures, thanks to the close cooperation with WP1. The compounds obtained will be tested in cellulo and demonstrated target engagement using the collaboration with WP 3.
 The project will focus on the identification of novel hit compounds following a fragment-screening approach. The optimisation of the best hit compounds will be done by monitoring both the in vitro activities on target and the cell-based activity. The frequency-of-resistance development of the most promising new compounds via the generation of resistant strains will be pursued.
The project will focus on the identification of novel hit compounds following a fragment-screening approach. The optimisation of the best hit compounds will be done by monitoring both the in vitro activities on target and the cell-based activity. The frequency-of-resistance development of the most promising new compounds via the generation of resistant strains will be pursued.
Our Team


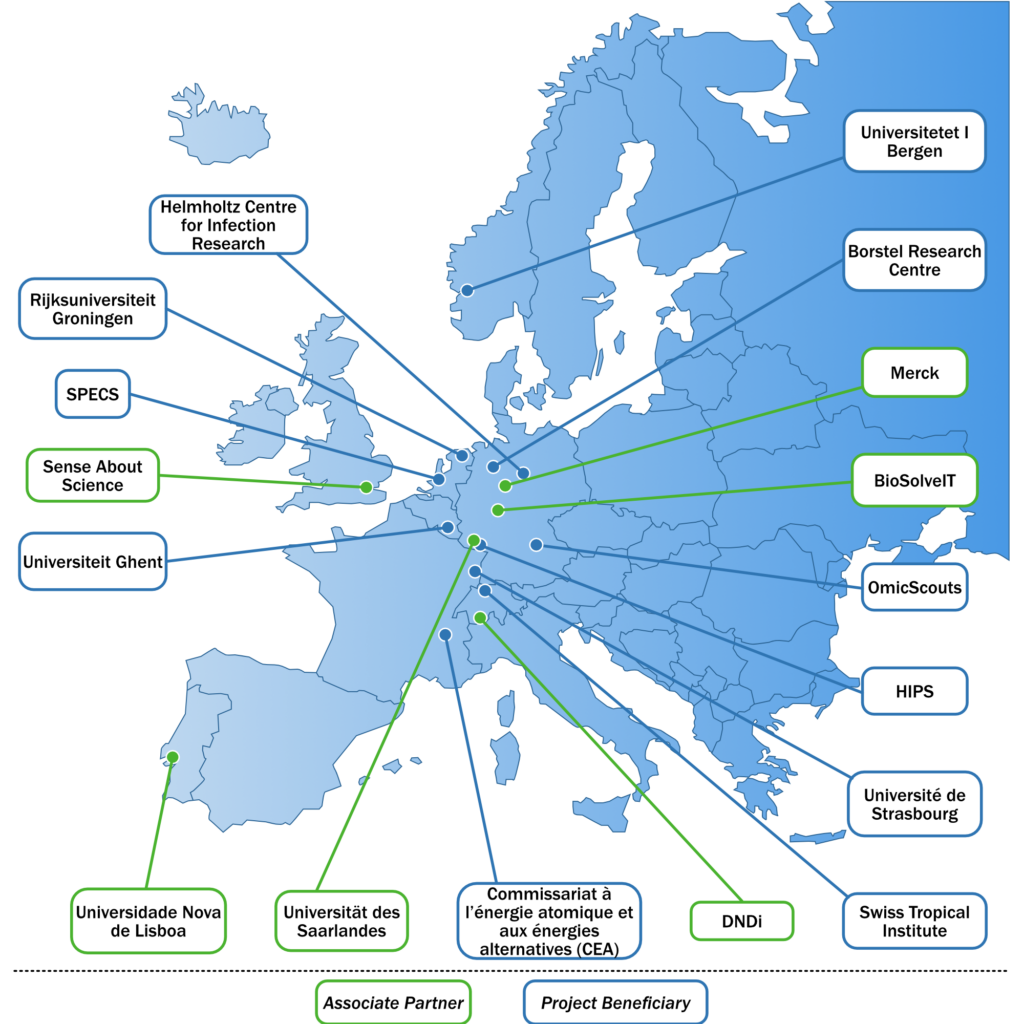
Project beneficiaries
NEWS
- MepAnti Drug Research Symposium: Young Minds, Big ImpactAnnouncement: the MepAnti team is organizing the MepAnti Drug Research Symposium: Young Minds, Big ImpactMore information about the symposium can be found here /*! elementor …
Continue reading "MepAnti Drug Research Symposium: Young Minds, Big Impact"
FUNDING

This project has received funding from the European Union's Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement No 860816.